
- Experimental metadata management
- Automated association of CRO files
- Automated infrastructure management
- nf-core pipeline support
- RStudio and Jupyter support
Your bioinformatician is running workflows in Colab and storing results on a laptop. Your metadata lives in a spreadsheet that one person understands. Setting up “real” compute infrastructure takes 2–6 months. Half a year before your first pipeline run.
I’ve watched this up close for years as the husband of a computational biologist. And as someone who’s spent 18 years building software infrastructure, I can confidently say that none of these problems are new. They’re solved problems, just not solved yet for small bio teams.
That’s why I built bioAF, an open-source computational biology automation framework.
One tool gives your team production-grade pipelines (Nextflow, nf-core), notebooks (Jupyter, RStudio), experiment tracking, and cost management all running on your own Google Cloud account. No vendor lock-in. No per-seat fees. No custom magic to untangle later.
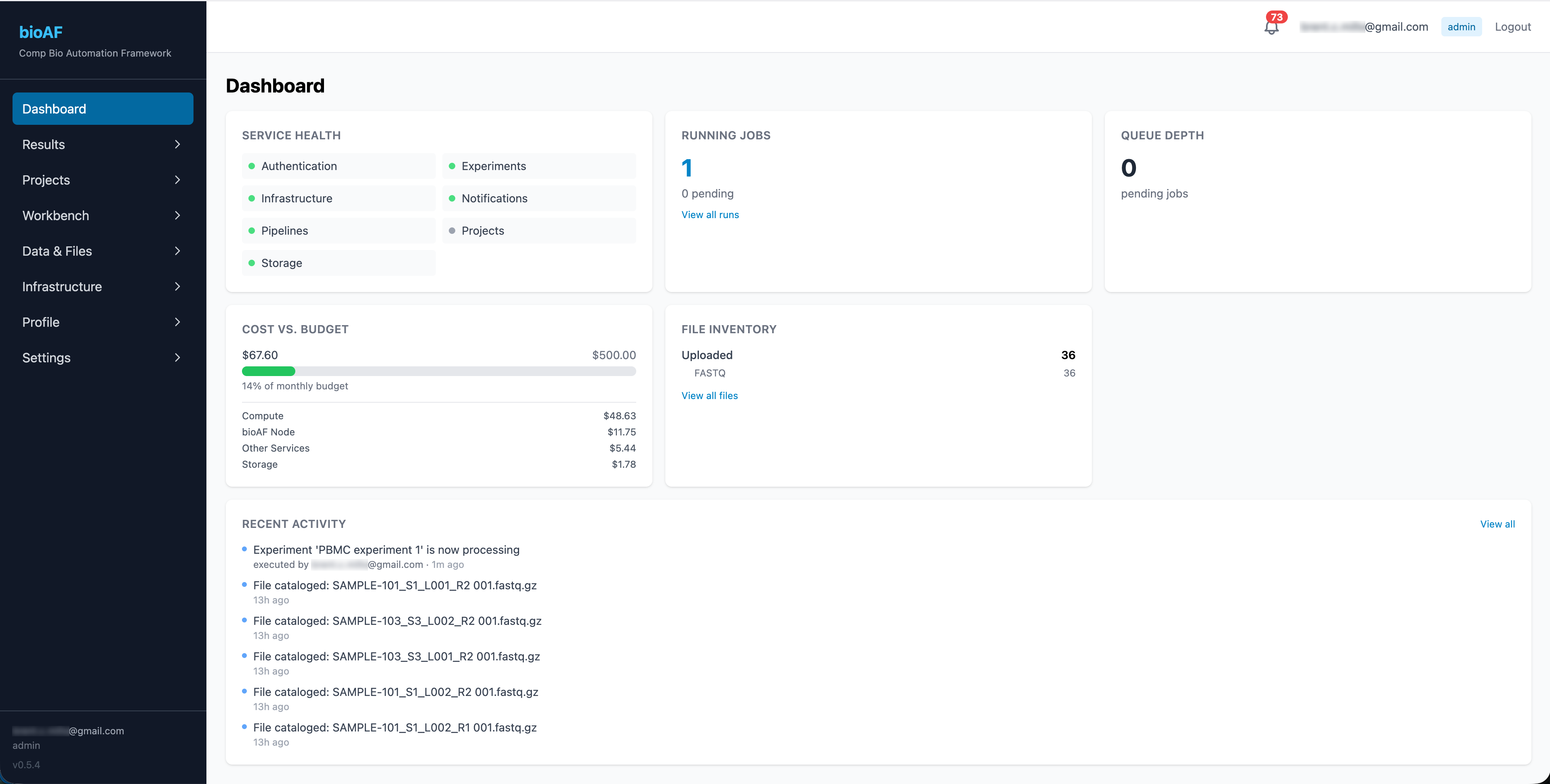
Bioinformaticians aren’t infrastructure engineers
Bioinformaticians and computational biologists are brilliant. But they aren’t infrastructure engineers or cloud architects. They don’t need to learn these skills.
bioAF doesn’t invent new tools. It wires together Kubernetes, Terraform, PostgreSQL, and GCS so your team never has to touch any of them directly. It offloads the infrastructure work and lets your team focus on the science.
The source is open. The tool works today. Check it out on bioAF.co or on Github.
Could this help your lab or company get to results faster? I’d love to hear what’s slowing you down.