Guide for Bench Scientists
This guide covers the day-to-day workflow for bench scientists using bioAF. Everything here happens in your web browser, no command line or coding required.
Registering an experiment
- Click New Experiment from the Experiments page
- Fill in the experiment metadata:
- Name and description
- Organism and tissue type
- Chemistry (e.g., 10x Chromium 3’ v3)
- Any relevant experimental conditions
- Click Create
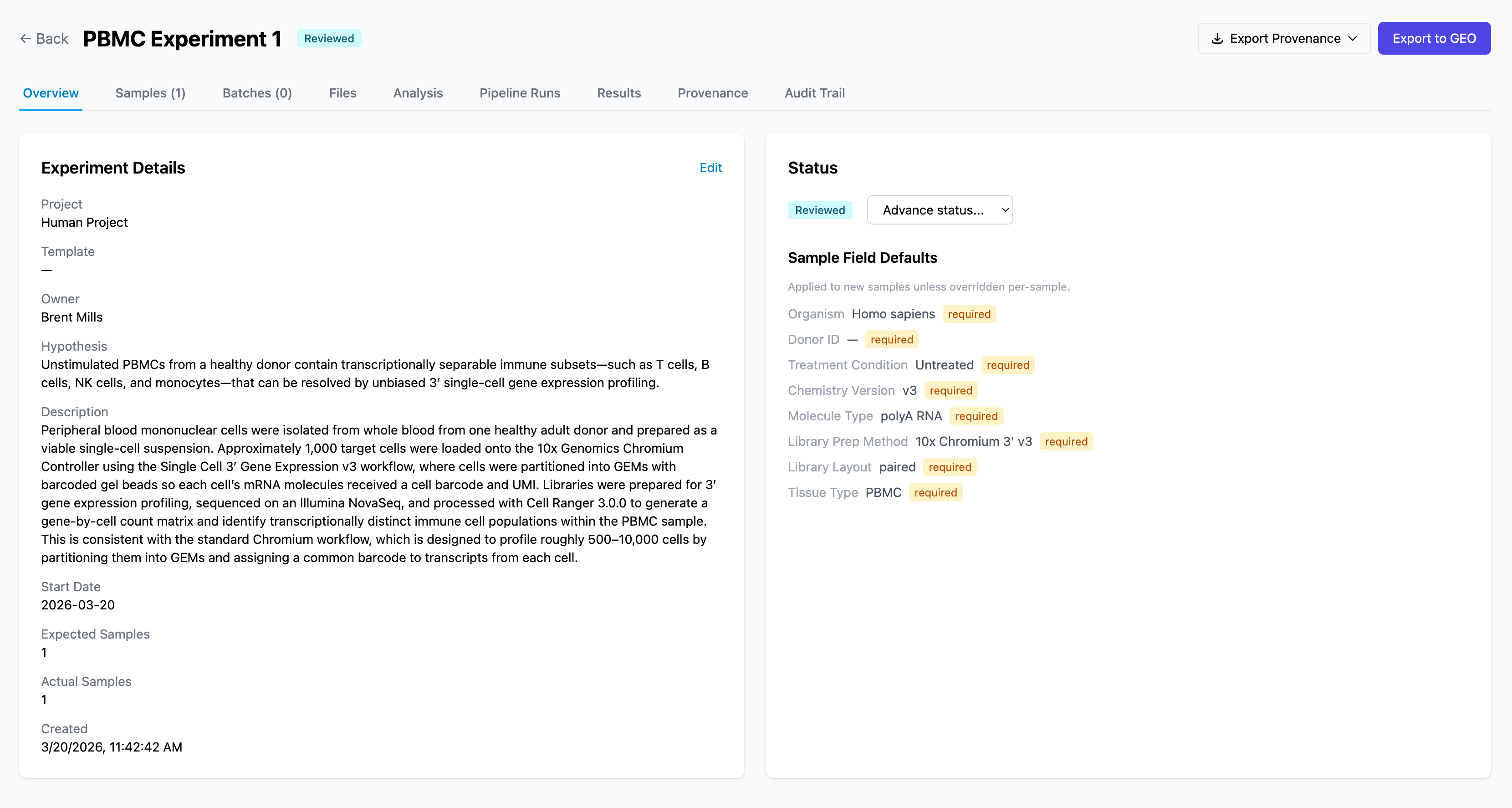
bioAF uses MINSEQE-compliant metadata standards so your experiments are publication-ready from the start.
What is MINSEQE?
Adding samples
- Open your experiment and click Add Samples
- For each sample, enter:
- Sample name and description
- Tissue type and cell type
- Treatment conditions
- Any custom metadata fields
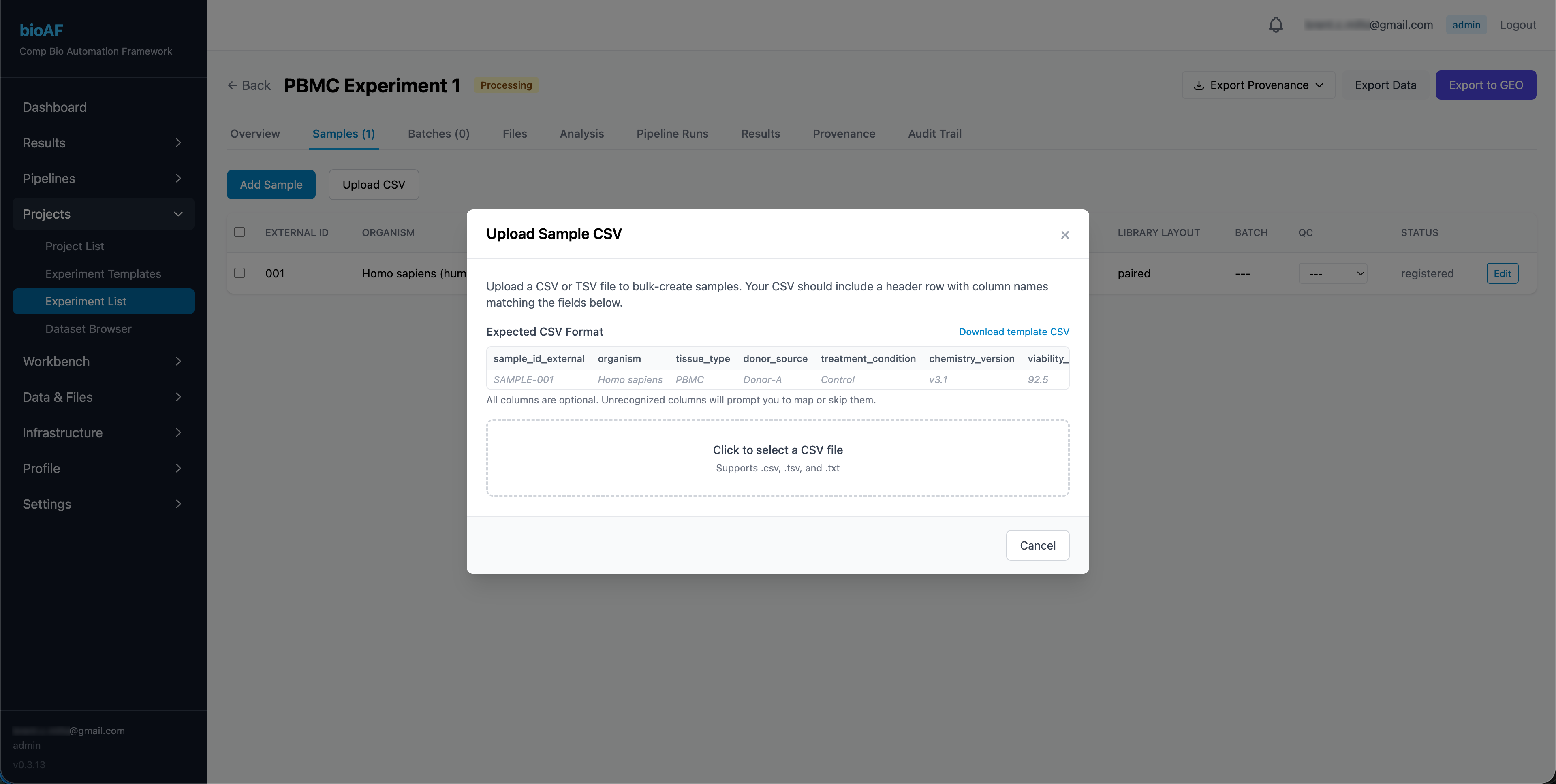
- You can add samples one at a time or batch upload from a spreadsheet
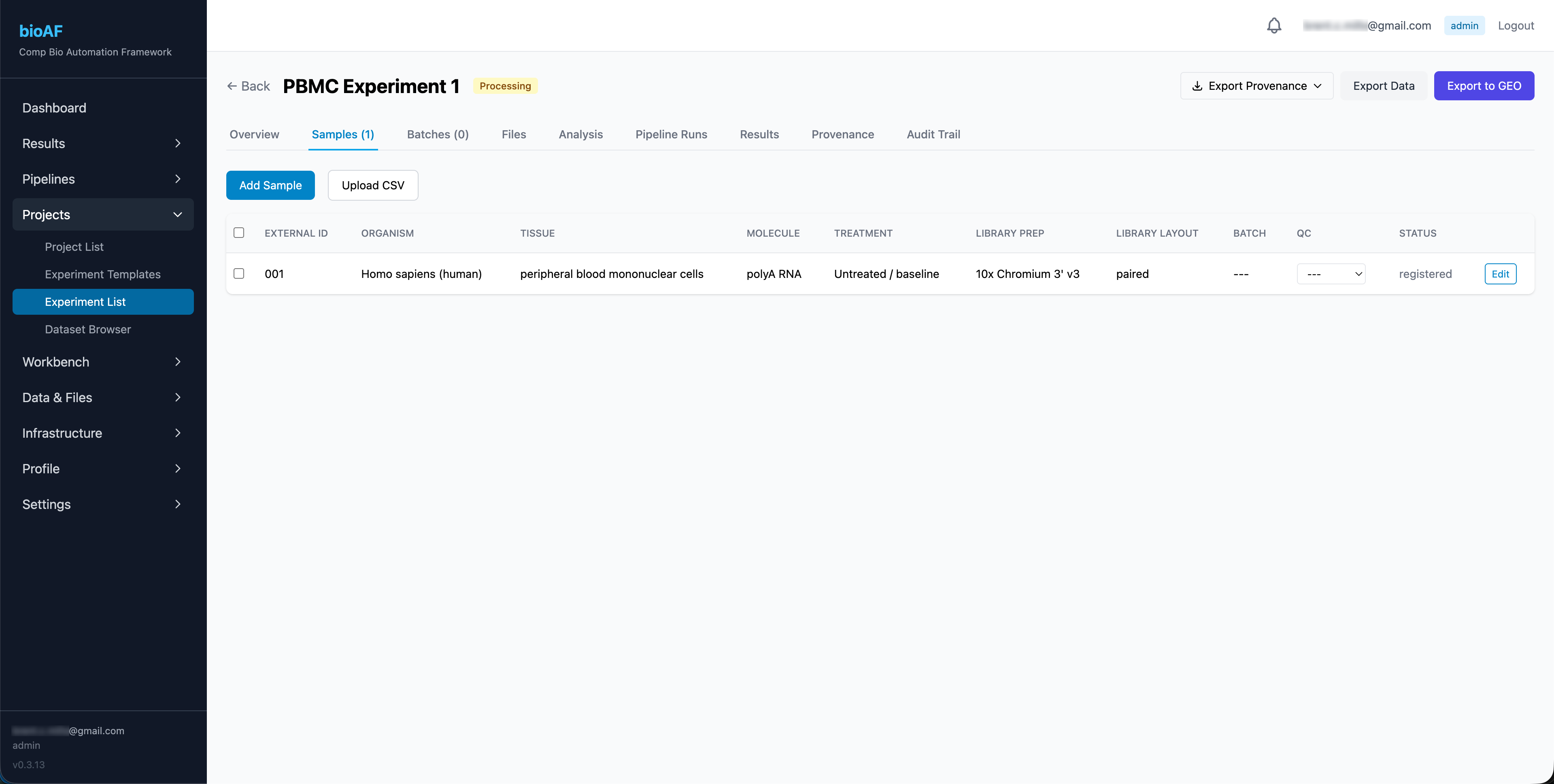
Uploading FASTQ files
- Navigate to your experiment’s Data tab
- Drag and drop your FASTQ files, or click Upload
- bioAF automatically associates files with the correct samples based on naming conventions
Can the sequencer upload automatically?
Viewing results
Once a pipeline has run on your experiment:
QC dashboards
Navigate to Results > QC on your experiment page. The QC dashboard shows quality metrics for your sequencing run, cell counts, read depth, mapping rates, and more.
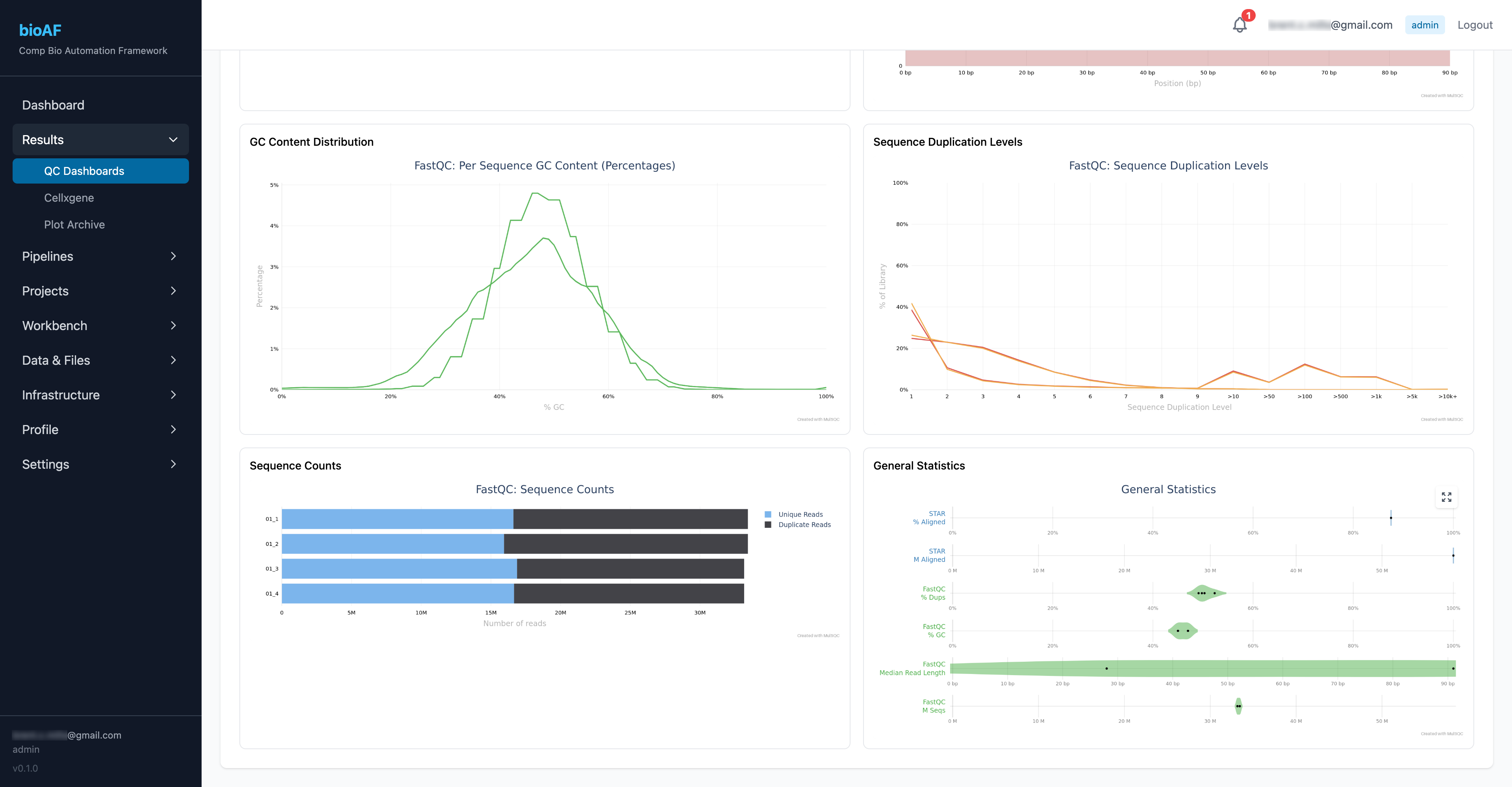
Interactive single-cell visualization
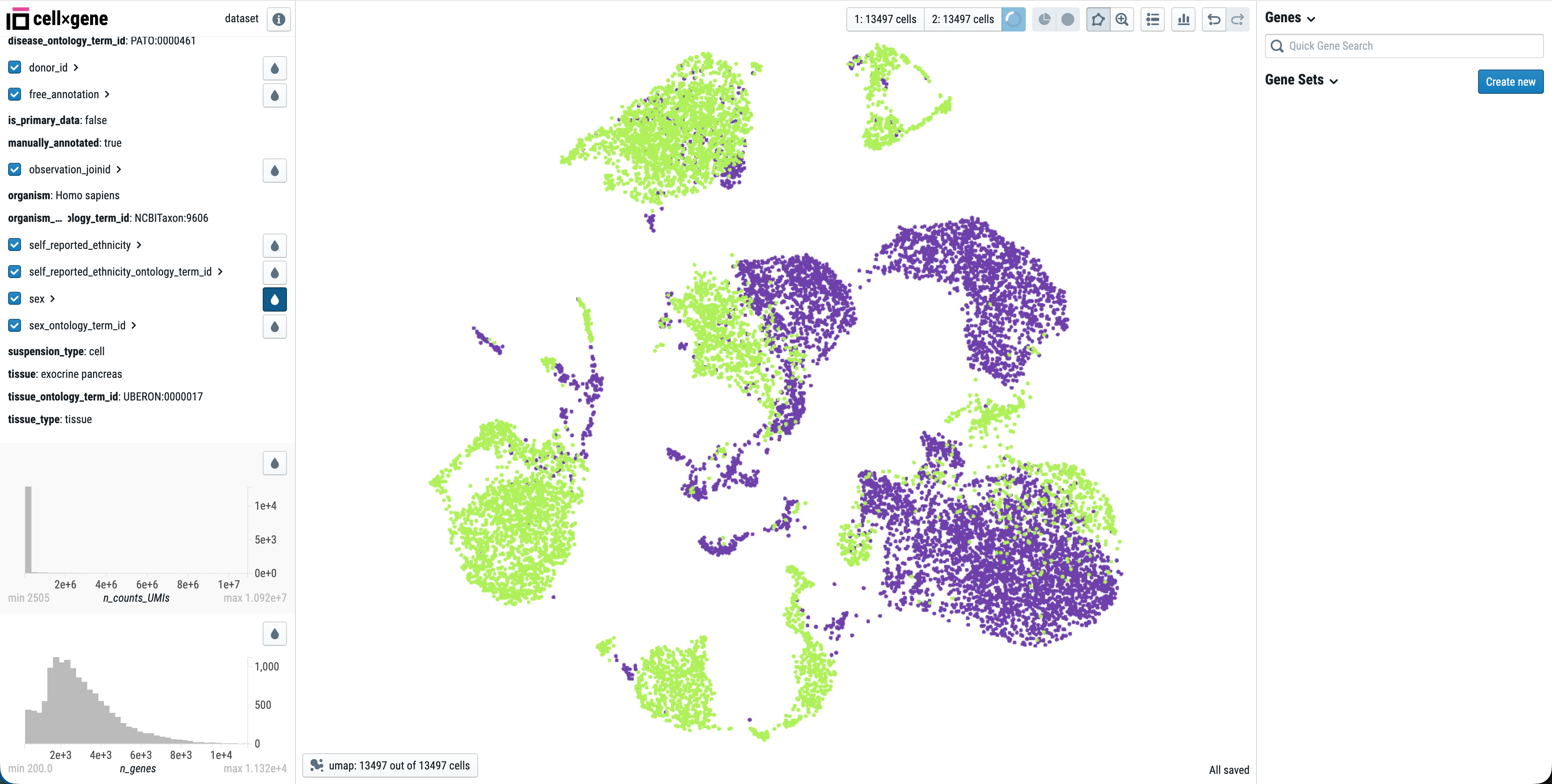
Click View in cellxgene to open an interactive explorer for your single-cell data. You can:
- View UMAP/t-SNE embeddings
- Color cells by gene expression, cluster, or metadata
- Select cell populations for comparison
- Export plots
Downloading results
From the Data tab, you can browse and download any output files, count matrices, processed h5ad files, plots, and reports.
Getting notifications
bioAF can notify you when:
- A pipeline finishes running on your experiment
- QC results are available
- Someone comments on your experiment
- Your experiment’s status changes
Check your notification preferences under your Profile settings.