Data Management
bioAF provides centralized data management with automatic organization and lifecycle policies.
File browser
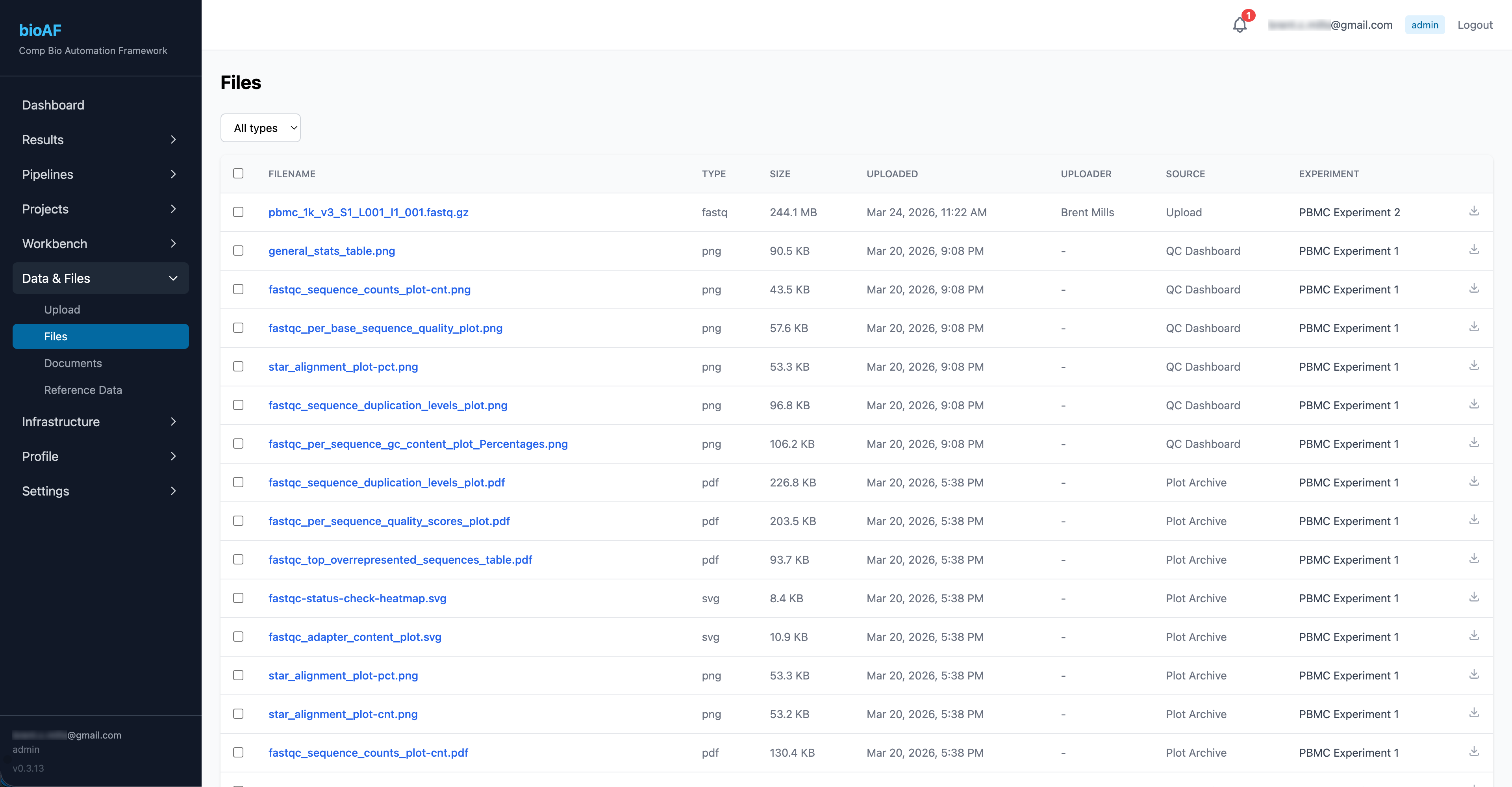
The Data & Files section gives you a unified view of all data on the platform:
- Browse by experiment, project, or file type
- See file metadata (size, upload date, source)
- Upload new files via drag-and-drop
- Download files individually or in bulk
Data lifecycle
bioAF organizes data across four storage tiers that map to the research workflow:
| Tier | Purpose | Retention |
|---|---|---|
| Ingest | Incoming files from sequencer or upload | Moved to Raw after processing |
| Raw | Original FASTQ files, untouched | Permanent |
| Working | Intermediate pipeline outputs | 30 days (configurable) |
| Results | Final outputs (h5ad, plots, QC reports) | Permanent |
Where is my data stored?
All data is stored in Google Cloud Storage (GCS) buckets in your own GCP project. You have full ownership and access, bioAF organizes and manages the data but never moves it outside your project.
Auto-ingest
If your sequencing core or instrument is on the same network, bioAF can automatically detect and import new FASTQ files as they’re produced. This uses Google Cloud Pub/Sub to watch for new files and trigger the import pipeline.
Dataset browser
The Datasets view provides a higher-level view of your data organized by experiment and analysis stage, making it easy to find specific results without navigating the raw file tree.