Guide for Bioinformaticians
This guide covers the computational biologist workflow in bioAF, running pipelines, working in notebooks, and managing compute environments.
Browsing and launching pipelines
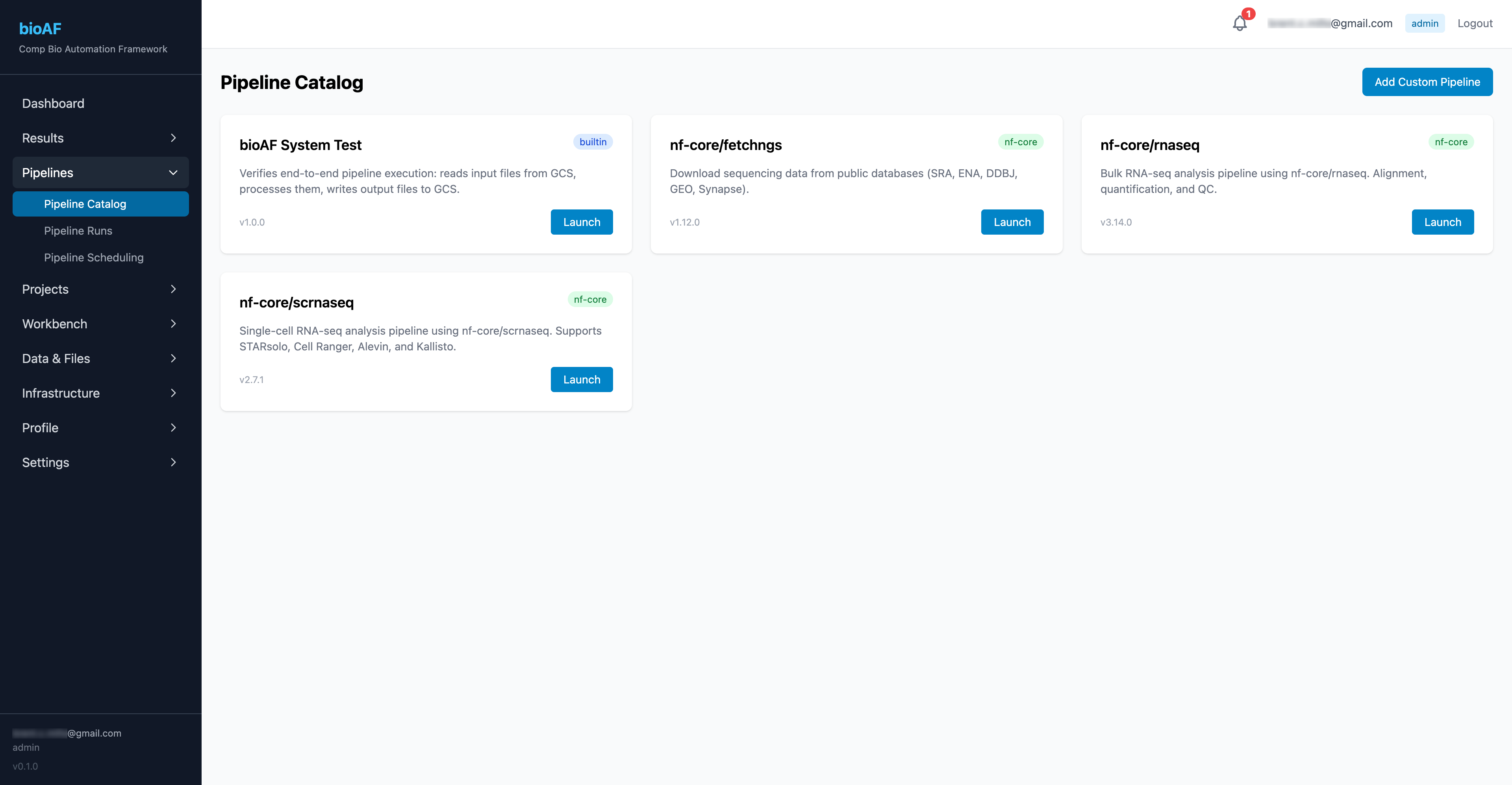
The pipeline catalog
Navigate to Pipelines to see the available workflows. bioAF ships with nf-core pipelines pre-configured:
- nf-core/scrnaseq: Single-cell RNA-seq (10x Chromium, Smart-seq2, etc.)
- nf-core/rnaseq: Bulk RNA-seq
- nf-core/atacseq: ATAC-seq
- And more, with the ability to add custom pipelines
Launching a pipeline run
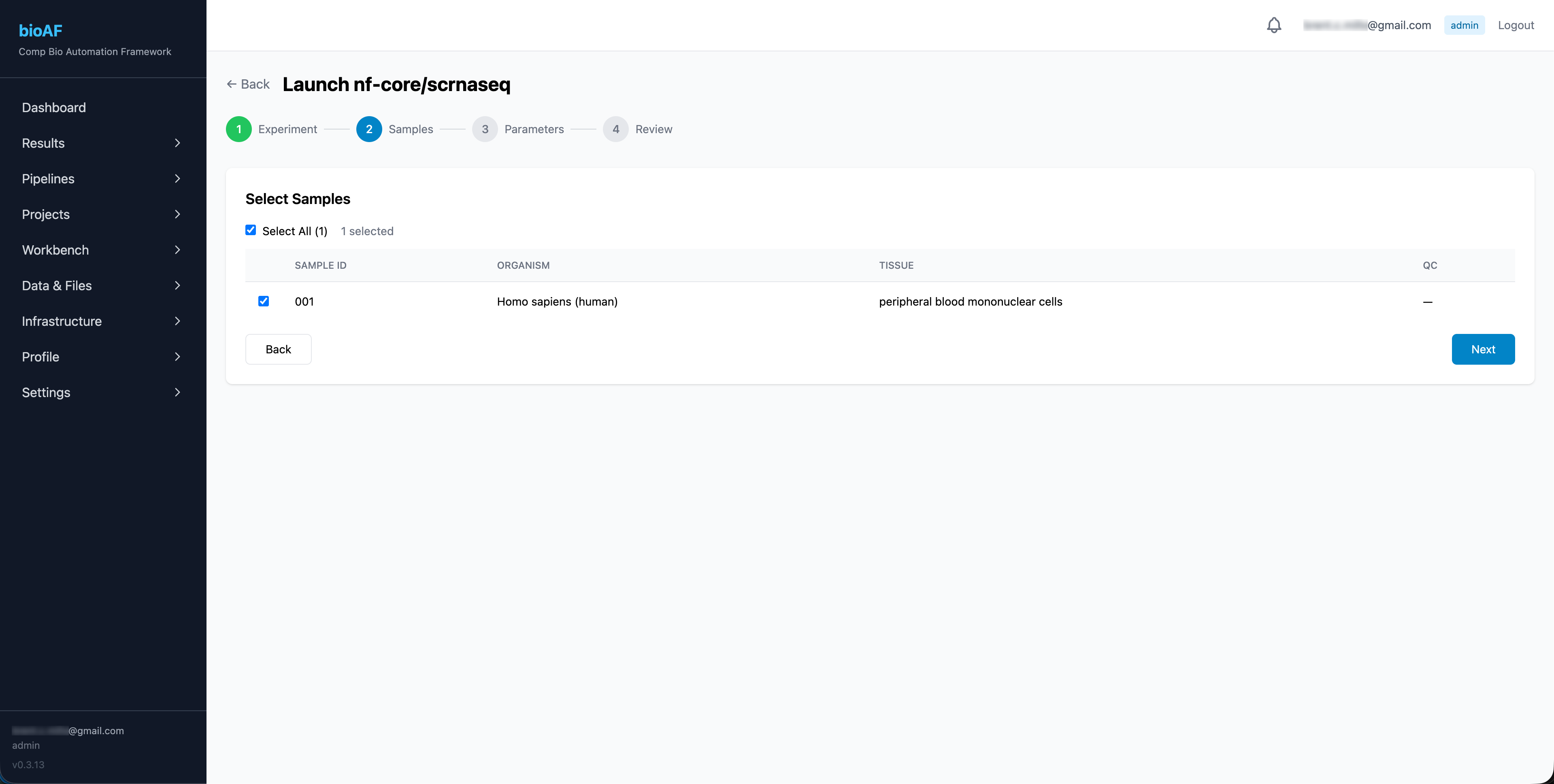
- Select a pipeline from the catalog
- Choose the experiment and samples to process
- Configure parameters (or use defaults)
- Set compute resources (CPU, memory)
- Click Launch
What is Nextflow?
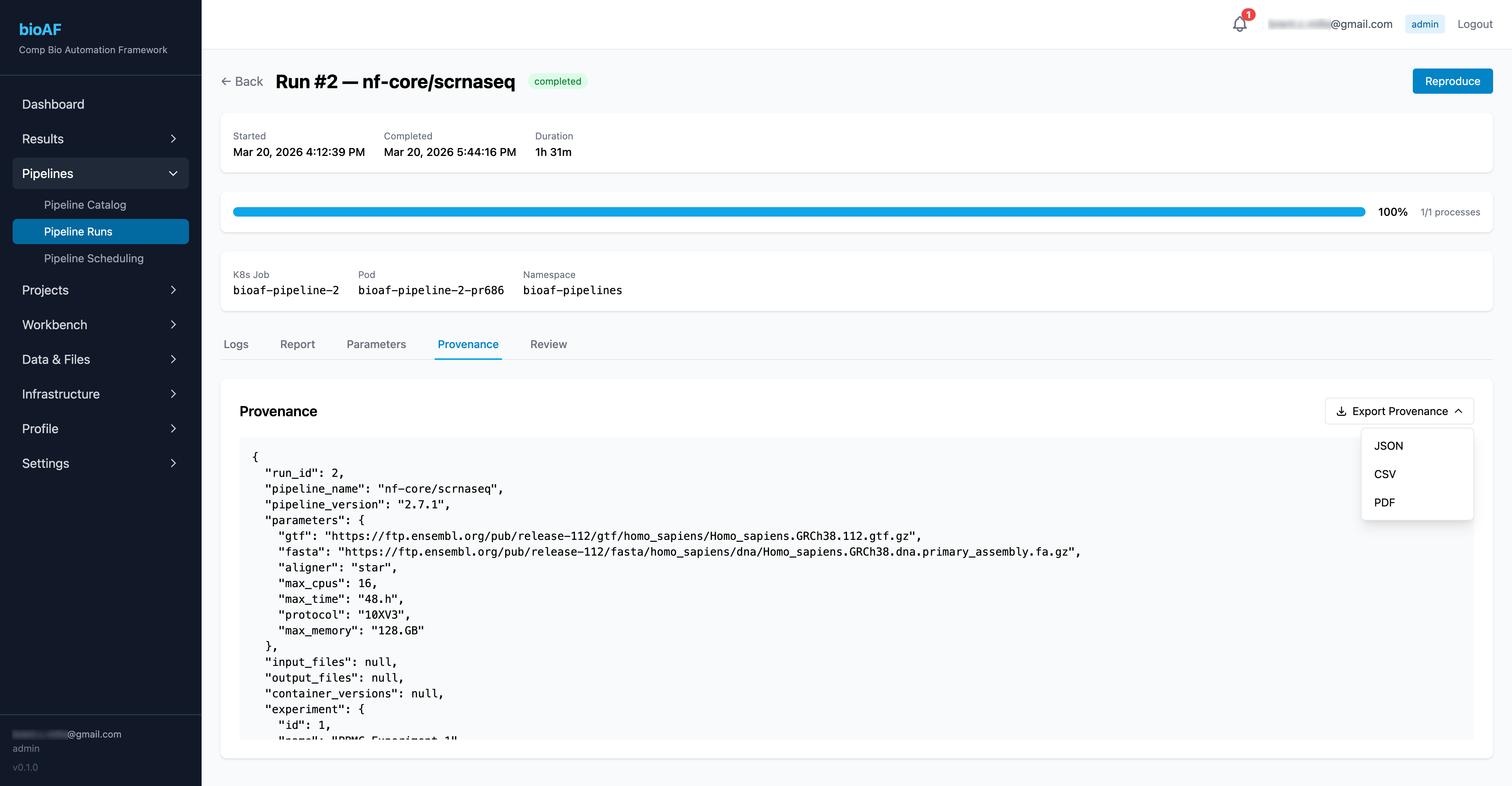
Monitoring pipeline runs
Once launched, you can track progress from the Runs page:
- Real-time status for each pipeline stage
- DAG (directed acyclic graph) visualization showing the workflow structure
- Resource usage and timing per stage
- Access to logs for troubleshooting
Working with notebooks
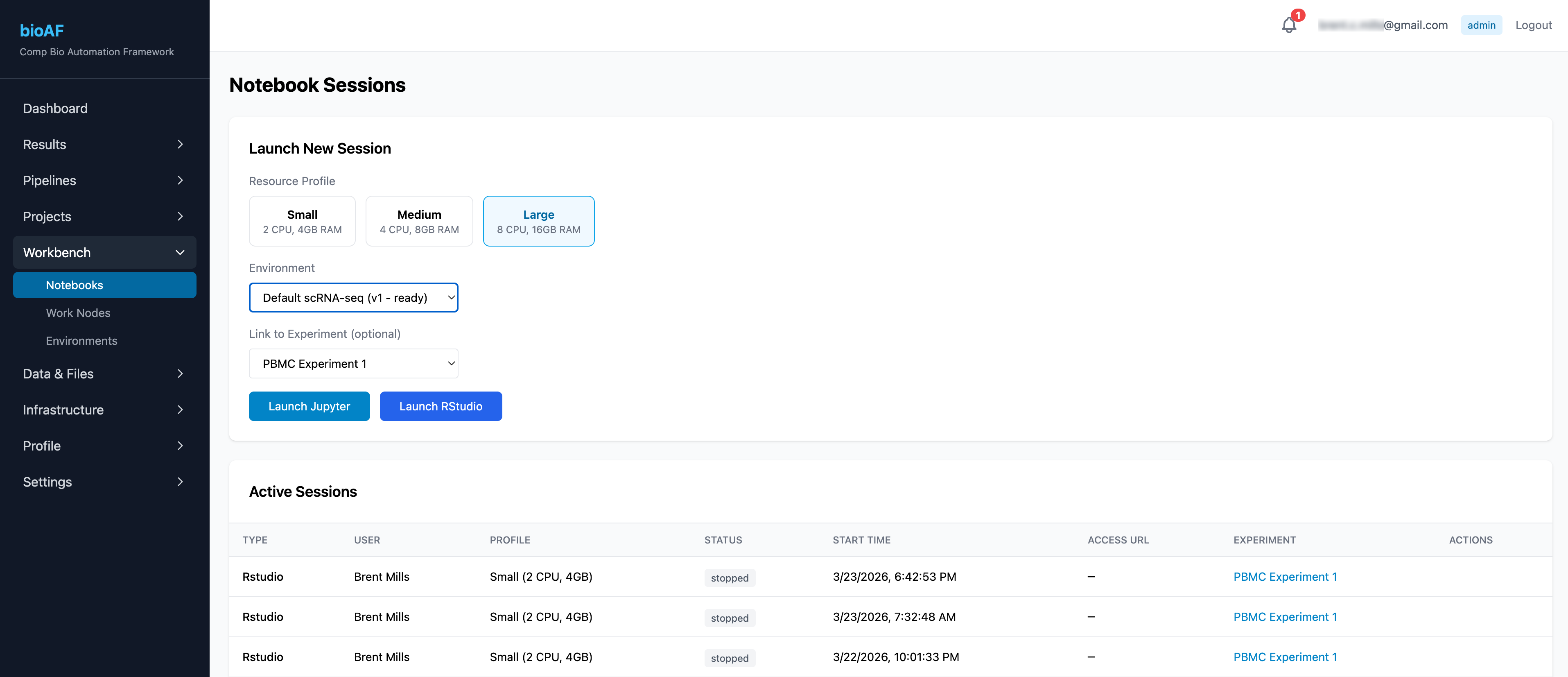
Launching a session
- Navigate to Notebook Sessions
- Click New Session
- Choose your environment:
- JupyterHub: For Python-based analysis
- RStudio: For R-based analysis
- Select a versioned environment (or use the default)
- Set resource limits and click Launch
Your session starts in about a minute. Click Open to connect directly in your browser.
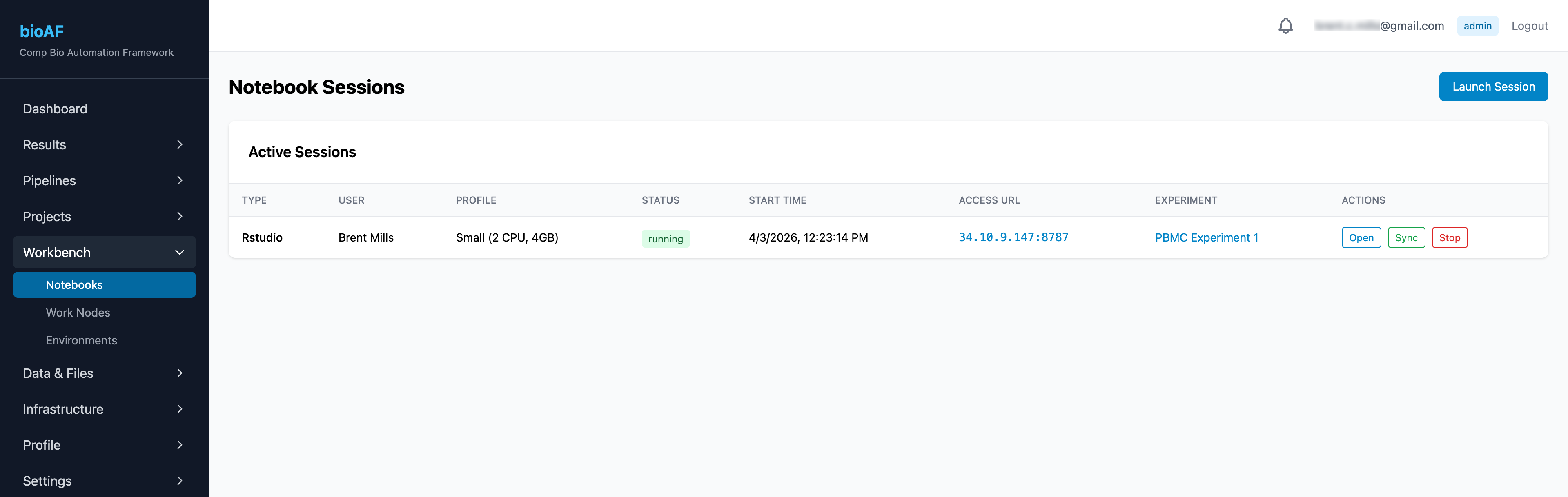
Sessions automatically stop after a configurable idle period to manage costs. Your work is saved, you can restart and pick up where you left off.
SSH access
For tasks that need a terminal, click the Terminal button on any running session or pipeline job. This opens a browser-based terminal directly into the running container.
Managing environments
What are environments?
Environments define the software available in your notebook sessions and pipeline runs, Python/R packages, system libraries, and tools. bioAF versions every environment so your analyses are reproducible.
Creating an environment
- Go to Environments > New Environment
- Provide a name and description
- Upload a Dockerfile or conda environment spec
- Click Build
bioAF builds the environment image and makes it available for notebook sessions.
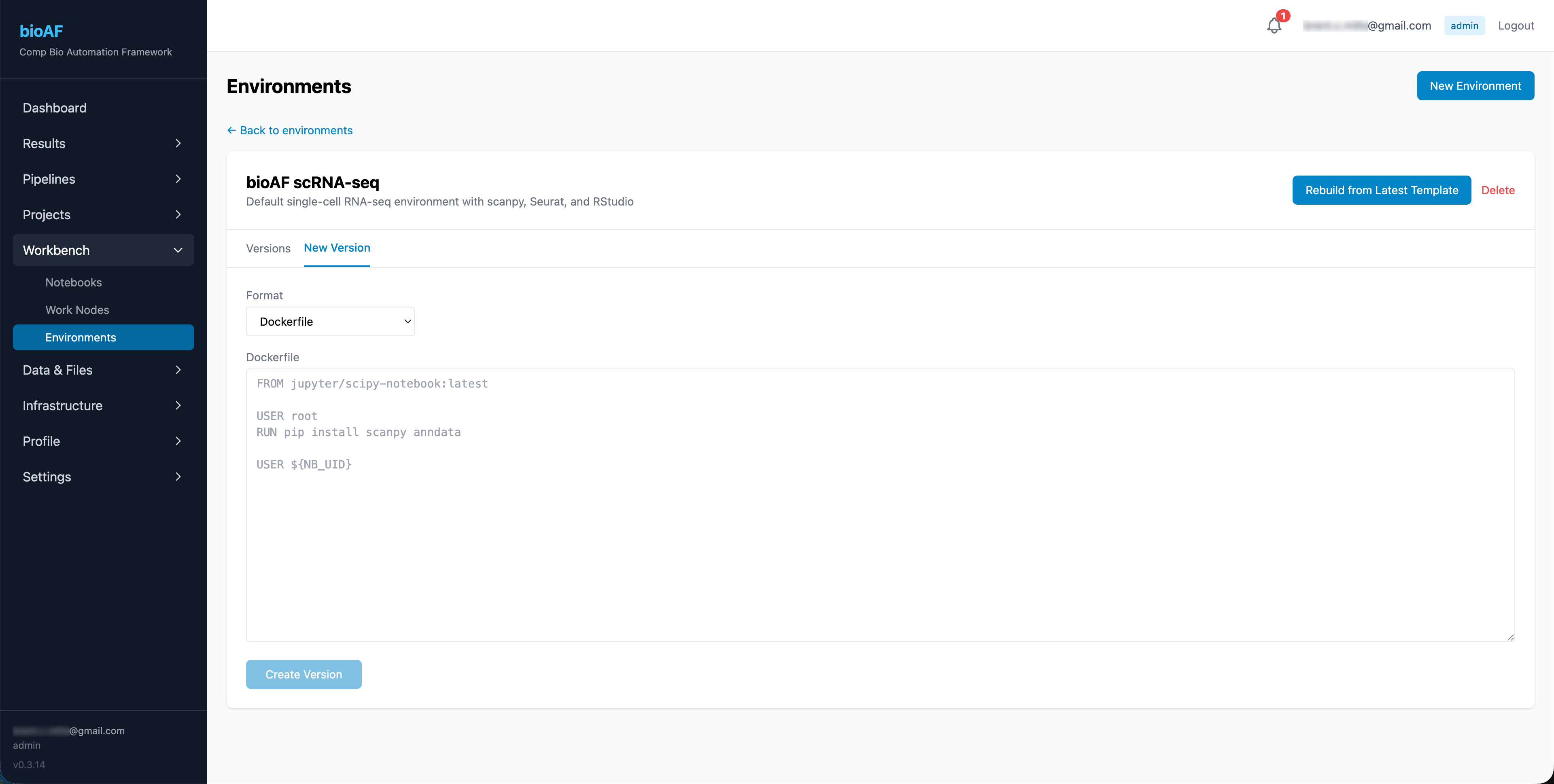
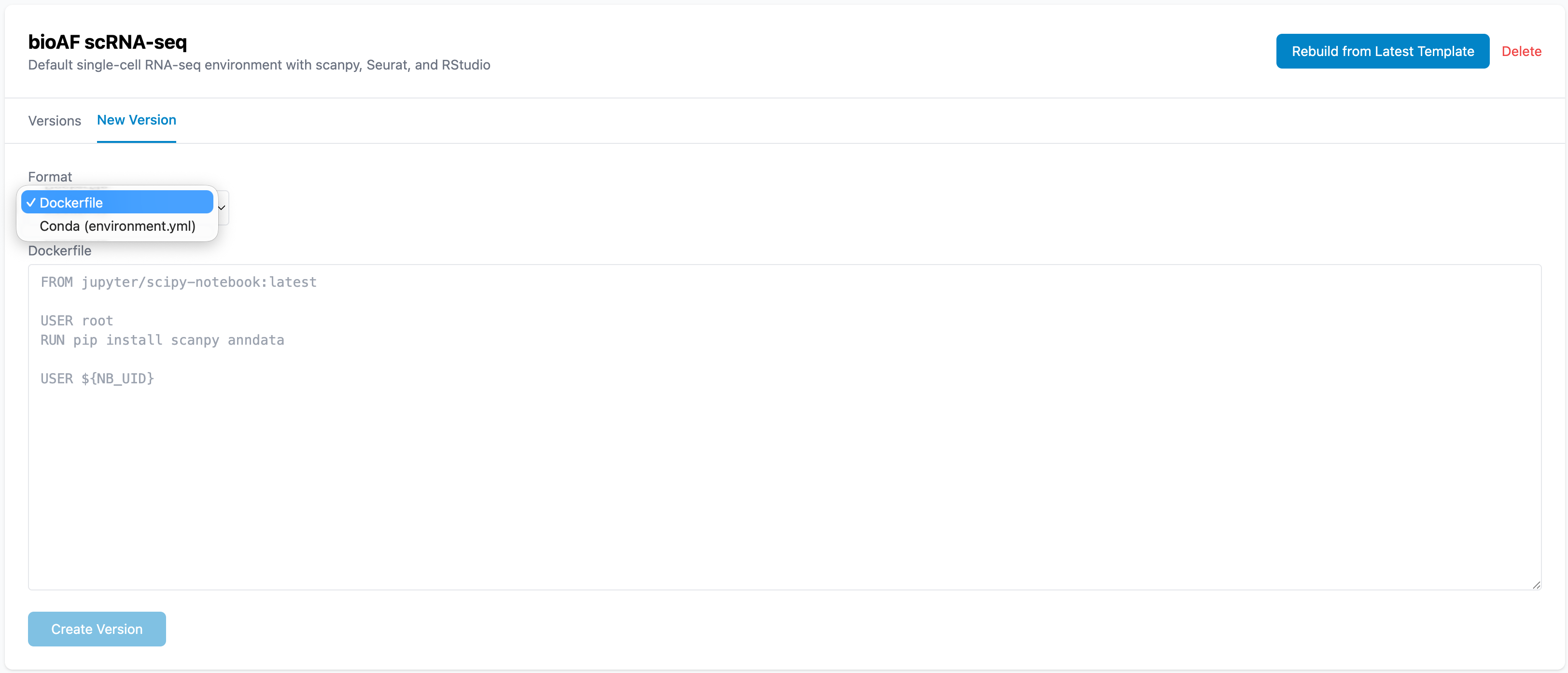
What is a Dockerfile?
Versioning
Every time you update an environment, bioAF creates a new version. Previous versions remain available, so you can always go back to the exact setup that produced a specific result.
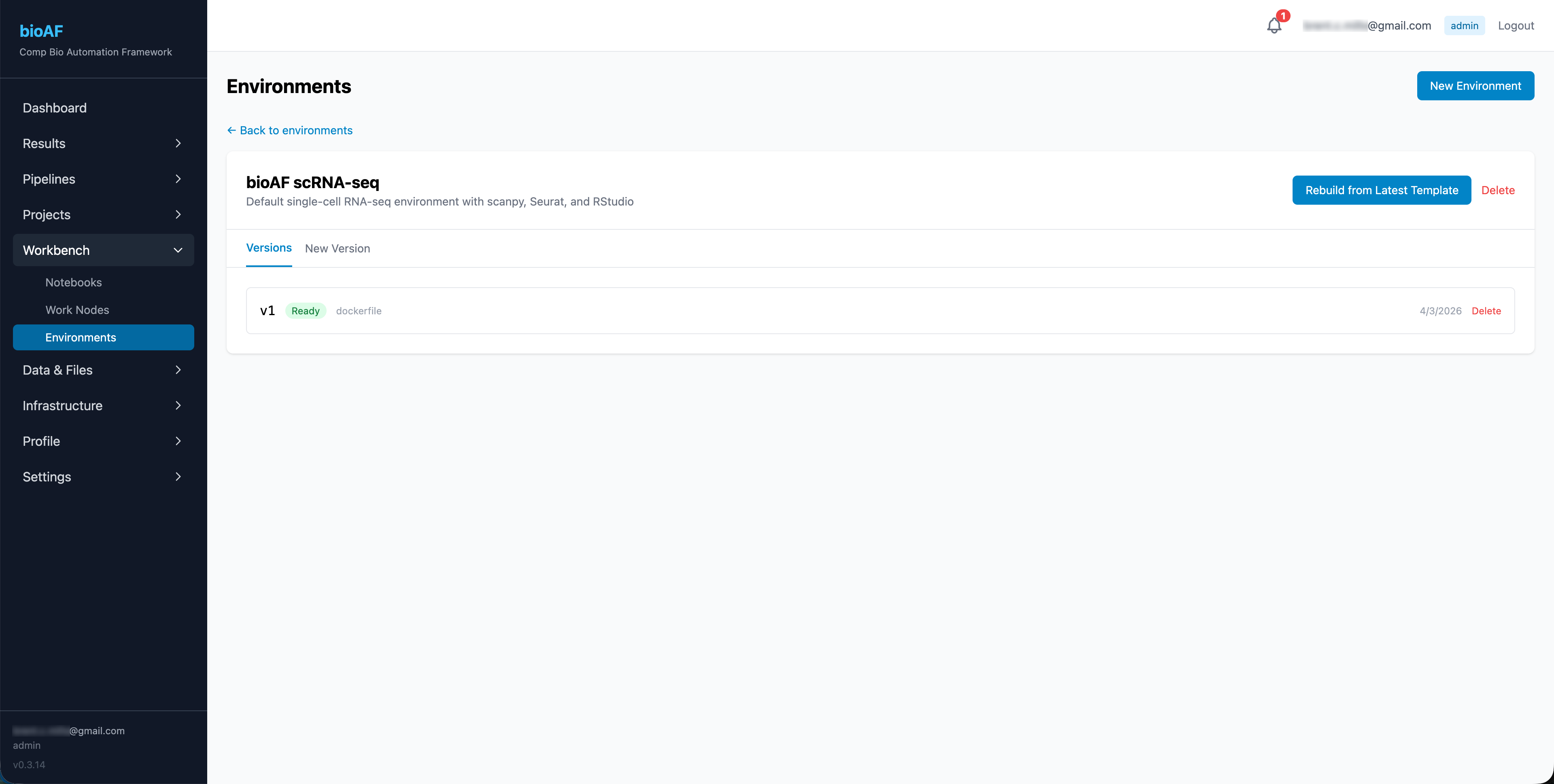
Accessing data and files
The Data & Files section lets you:
- Browse files by experiment or project
- Upload new files
- Download results, count matrices, and reports
- See file metadata (size, upload date, associated experiment)
Adding custom pipelines
If you have your own Nextflow workflows, you can add them to the pipeline catalog. Navigate to Pipelines > Add Pipeline and provide:
- Pipeline name and description
- Git repository URL
- Default parameters
- Resource recommendations