Pipeline Engine
bioAF includes a built-in pipeline execution engine that lets you run bioinformatics workflows without managing compute infrastructure.
Pipeline catalog
bioAF ships with pre-configured nf-core pipelines:
- nf-core/scrnaseq: Single-cell RNA-seq analysis
- nf-core/rnaseq: Bulk RNA-seq quantification
- nf-core/atacseq: ATAC-seq peak calling and analysis
- Custom pipelines can be added by providing a Git repository URL
What is nf-core?
nf-core is a community effort to collect, curate, and maintain high-quality Nextflow pipelines for bioinformatics. These pipelines are peer-reviewed, well-tested, and follow best practices. bioAF makes them available with one-click launching.
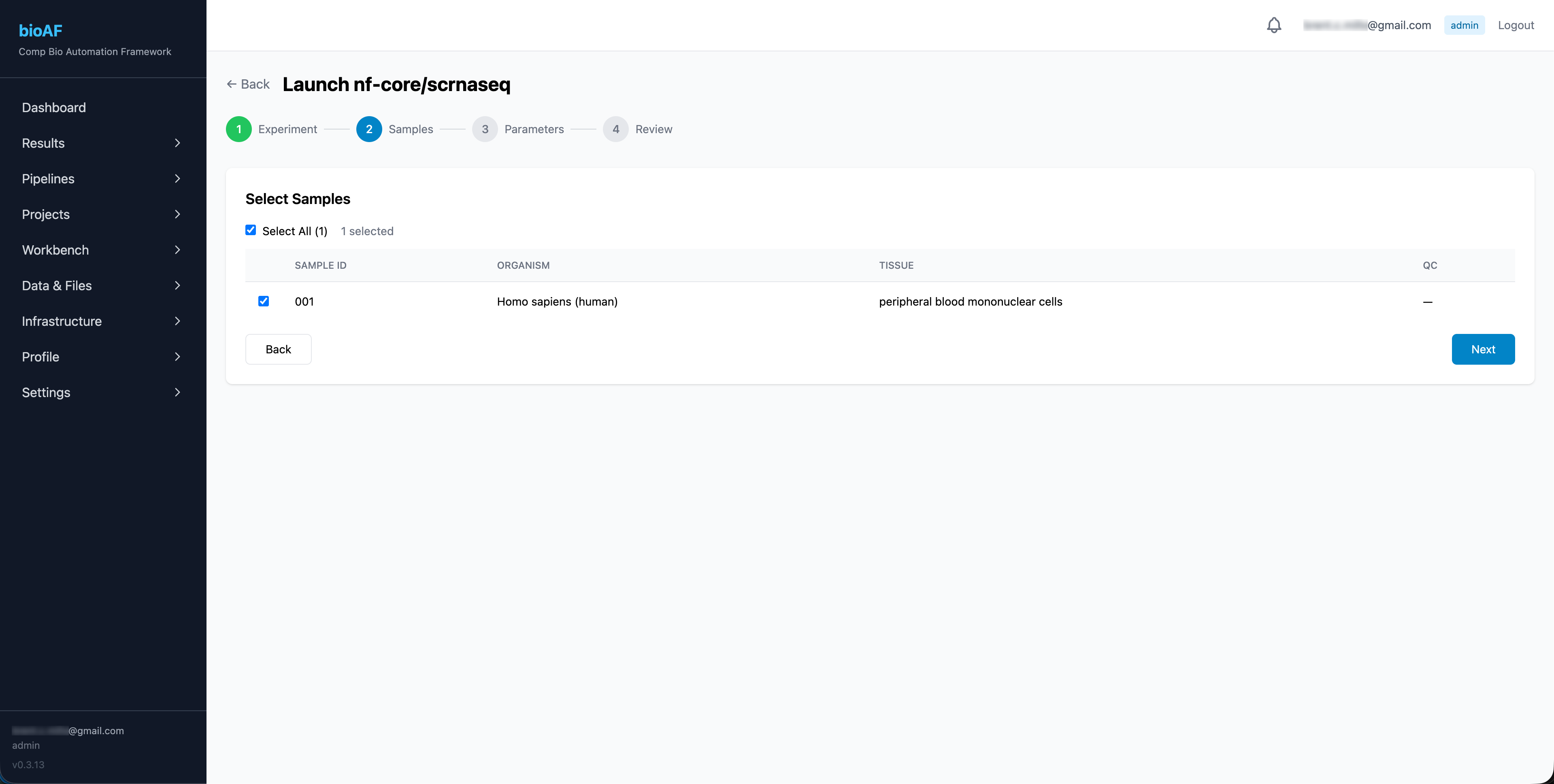
Launching a run
Select a pipeline, choose your experiment and samples, configure any parameter overrides, and launch. bioAF handles:
- Scheduling compute resources
- Staging input data
- Executing the workflow
- Collecting outputs to the results bucket
- Notifying you when it’s done
Real-time monitoring
Every running pipeline shows:
- Stage-by-stage status: See which steps are complete, running, or queued
- DAG visualization: A visual graph of the workflow structure
- Resource usage: CPU, memory, and timing per stage
- Live logs: Stream logs from any running stage
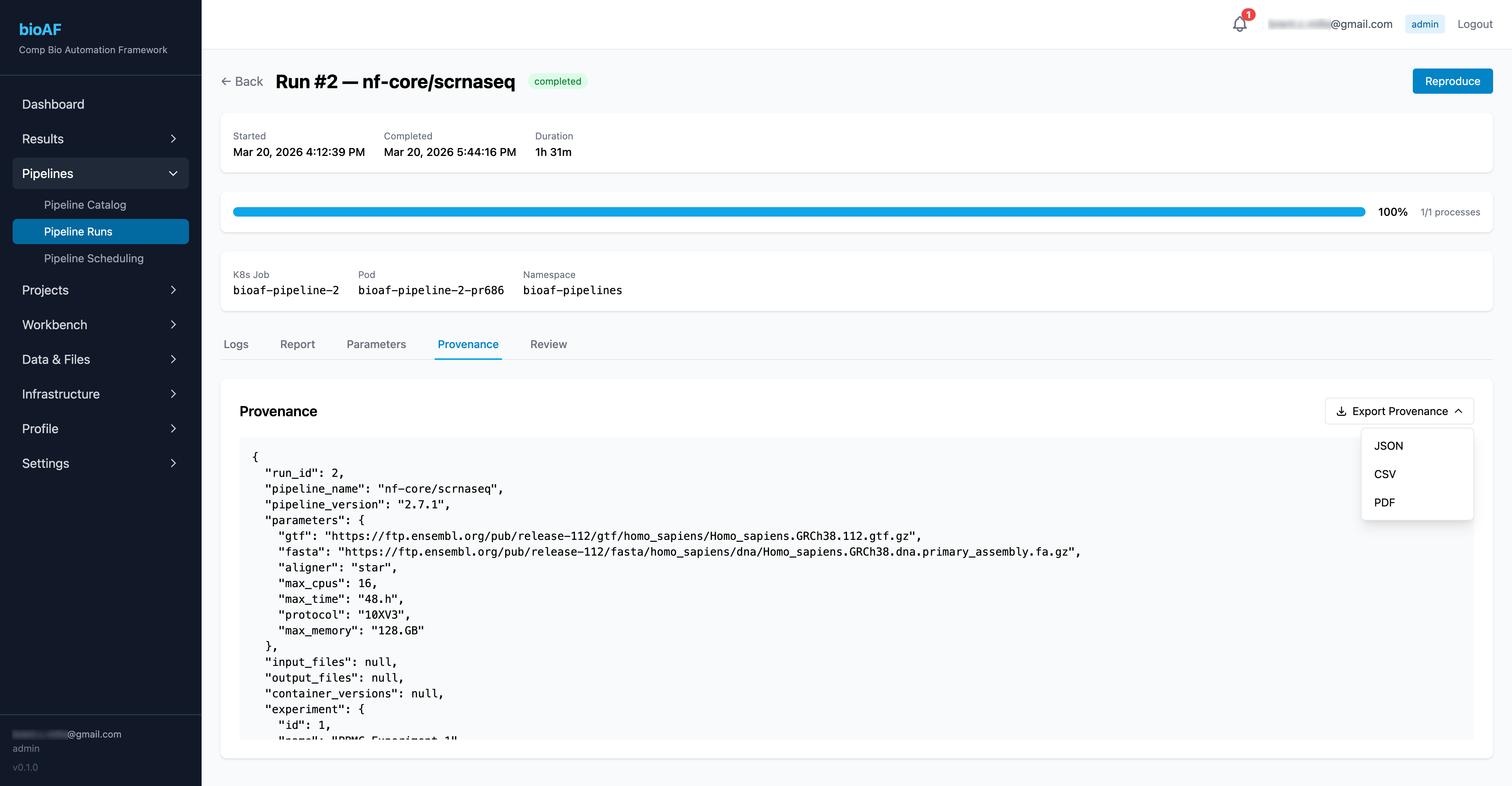
Outputs and results
When a pipeline completes, outputs are automatically:
- Stored in the results bucket
- Linked to the originating experiment and samples
- Available for QC dashboards and visualization
- Recorded in the audit log with full parameter provenance